Facilities
We host world-class, end-to-end facilities to rapidly design, scale and de-risk sustainable protein innovations from lab to market.
Cell engineering
Our AI-powered biofoundry automates pathway design, DNA assembly and high-throughput screening to generate optimised strains in days.
A droplet-microfluidic CRISPR platform with on-chip phenotyping plus real-time multi-omics dashboards and machine-learning slash fermentation optimisation from months to weeks.
Core Research Areas
- In-silico design – oligo synthesis: ML-powered pathway modelling feeds a DNA Script SYNTAX printer for overnight, on-site oligo production.
- Construct build – assembly: Labcyte Echo 525/550 acoustic dispensers and Biomek i7 robotics assemble thousands of plasmids daily in nanolitre volumes.
- Genome editing: A droplet-microfluidic array delivers multiplex CRISPR edits and electroporation at >10³ strains min-¹.
- HT screening & analytics: PIXL colony picker, iSeq100/MinION sequencing and Clariostar plate readers stream genotype-phenotype data to live dashboards.
- World class biofoundry access: London, DTU Biosustain & National University of Singapore
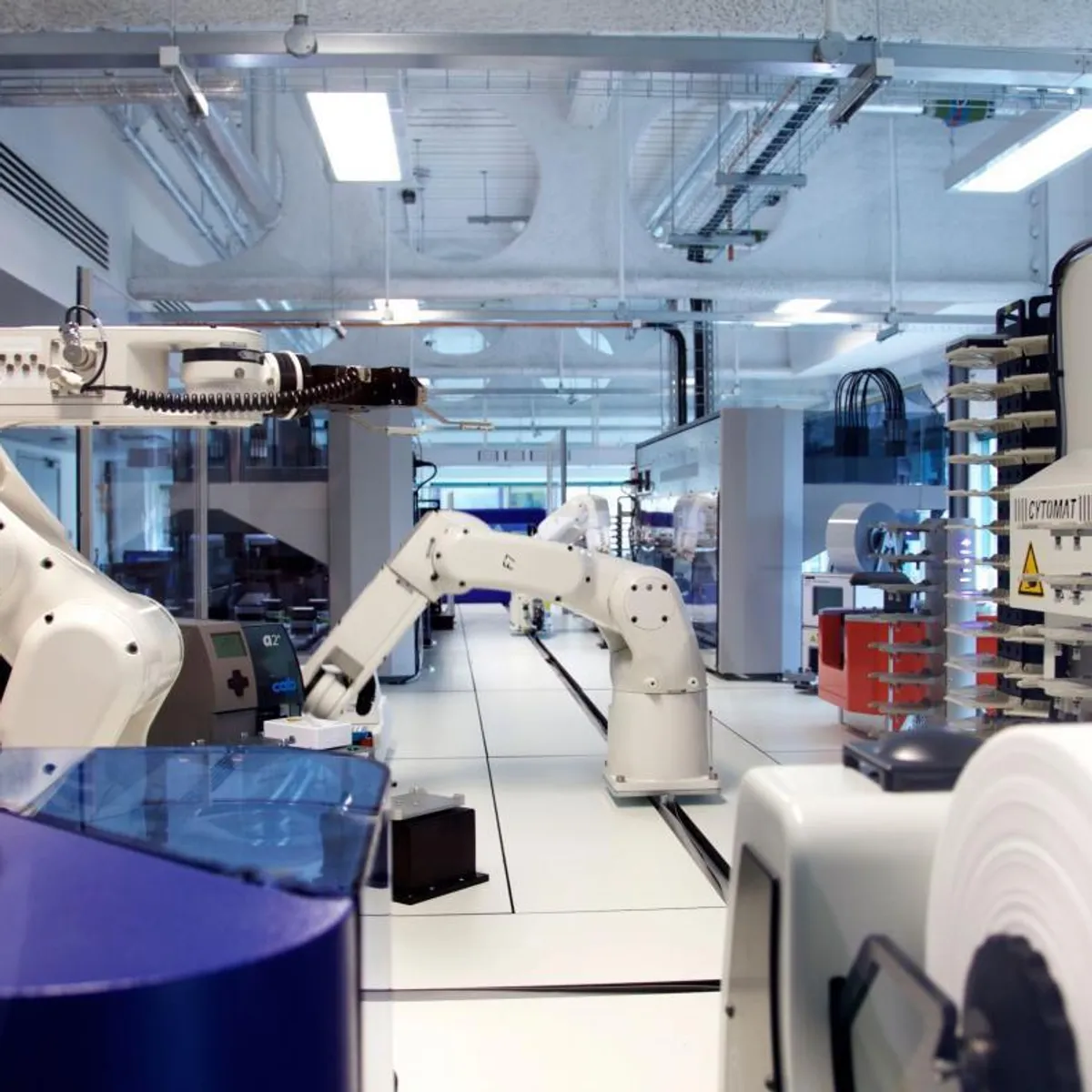
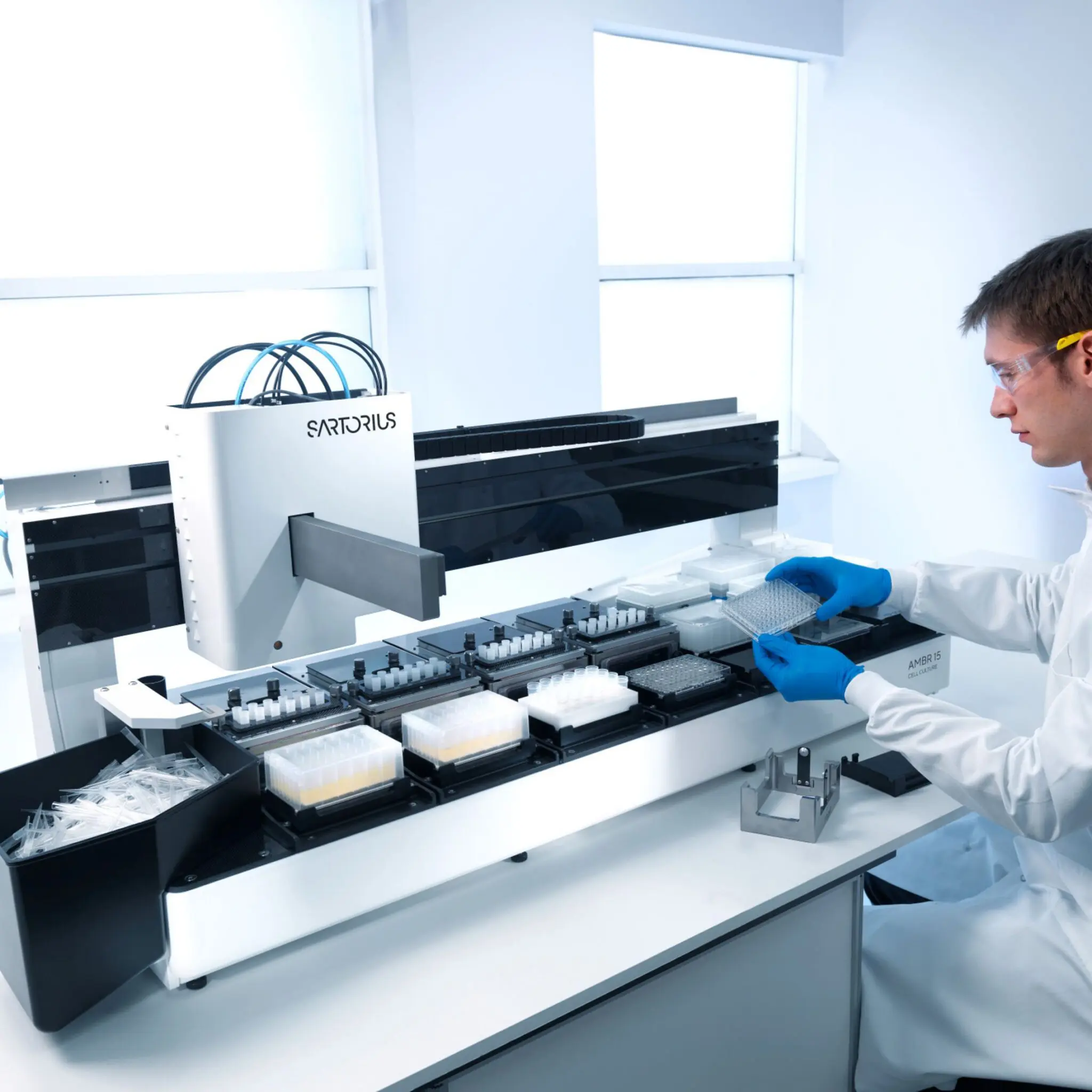


Scale UP – Microbial
Microbial & mammalian growth: our distributed bioprocess platform links ambr® 250 high-throughput microscale reactors with 300 L stainless-steel pilots and food-grade suites, letting us intensify and transfer processes across nine partner sites with one data backbone.
Inline Raman & capacitance into large data models, accelerating seed-to-pilot optimisation.
Core Research Areas
- Seed train & microscale DoE – 24-/48-parallel ambr® 250 HT reactors for microbial and perfusion/fed-batch screening (Imperial, Kent).
- Lab-scale optimisation (1–5 L) – Sartorius/Applikon STRs with inline Raman + dielectric probes in Imperial’s BioFlo suite.
- Mid-scale validation (20–100 L) – single-use and stainless STRs up to 100 L fully-functional pilot plant.
- Pilot fermentation (100–300 L) – food-grade 1-300 L suite at and 300 L pilot plant with U-loop fermenter for high-oxygen transfer rate processes.
- Downstream processing–Tangential flow filtration at scale to continuous centrifugation and chromatography there are a range of equipment available to suit your needs.
Scale UP – Plant
From sterile tissue culture to multi-hectare field trials and pilot biorefineries—is powered by a network of controlled-environment, agronomy and processing assets that spans the UK.
This integrated pipeline lets us take a newly edited lines from petri-dish to protein-rich powder in a single data backbone, collapsing varietal development and feedstock qualification timelines by an order of magnitude.
Core Research Areas
- In-vitro initiation: Class I/II aseptic tissue-culture and bioprinting suites generate transgenic or CRISPR-edited callus lines ready for rapid propagation.
- Controlled-environment phenotyping: >6,000 m² LED-equipped glasshouses and growth rooms run high-throughput DoE screens on a wide range crops.
- Greenhouse-to-hub transition: The Greentech Hub at NIAB East Malling and the Medway Food Innovation Centre provide climate-controlled propagation plus post-harvest testing.
- Multi-site field trials: Rothamsted’s 800 ha farm network (330 ha Harpenden core, 120 ha Brooms Barn, 350 ha North Wyke) delivers multi-soil, GM/GE-ready plots for yield and stress validation.
- Primary biomass processing: AberInnovation’s 100–1,000 kg h-¹ juicing and steam-explosion lines convert harvested biomass into fibre-free press-cake feedstocks for further fractionation.
- Pilot biorefining & fractionation: Food-grade downstream facilities (50–2,000 L h-¹ size-fractionation, spray-drying, membrane separation), complemented by novel seaweed and pulse protein extraction research, yield spray-dried concentrates ready for formulation.
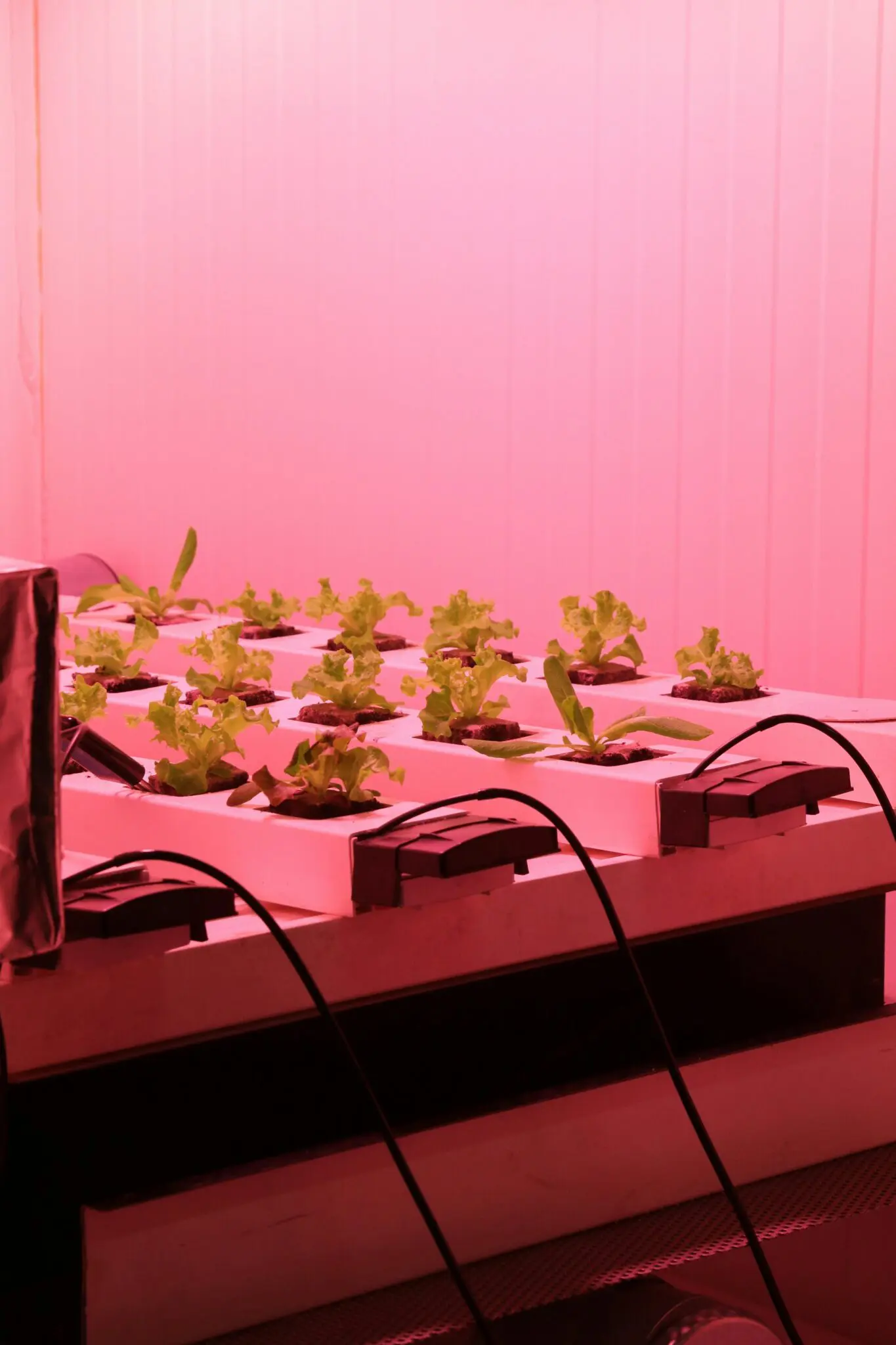
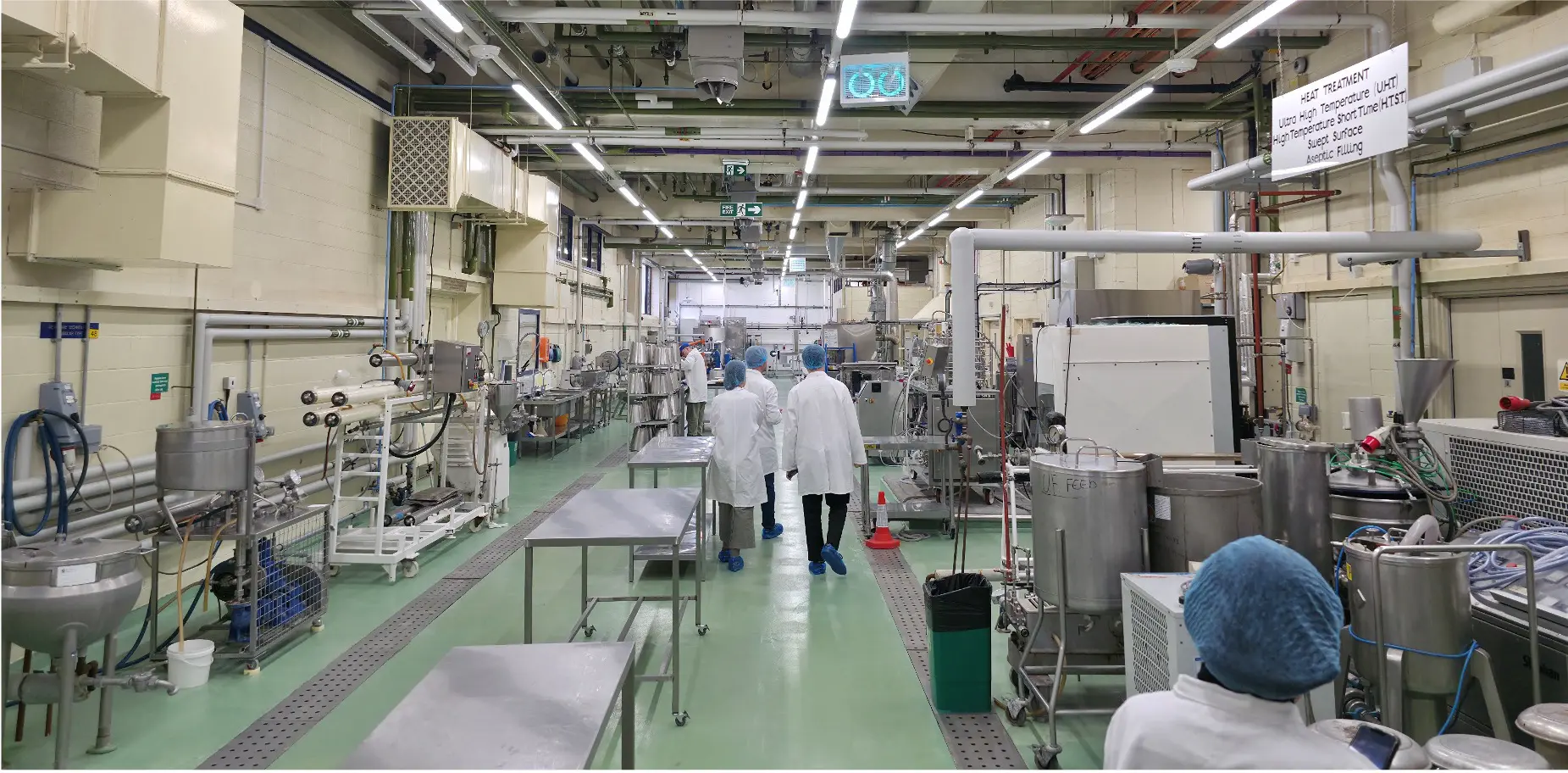


Food Processing Technologies
Food processing & testing—from raw ingredient to consumer-ready prototype—is driven by a unified network of pilot-scale equipment, sensory science suites and regulatory-grade analytical labs that accelerates product iteration and de-risks commercial launch.
Inline PAT feeds a secure digital-twin backbone, so process tweaks made on a 5 kg h-¹ bench line translate seamlessly to 100 kg h-¹ pilots, cutting optimisation timelines from quarters to weeks.
Core Research Areas
- Formulate, Test, Taste – Repeat – GMP compliant facilities to accelerate your product development with safe scalable processes and sensory testing ISO-compliant booths with trained panels.
- Ingredient pre-conditioning – precision milling, fractionation and cold-press oil extraction lines (≤ 200 kg h-¹) generate flours, protein concentrates and lipid fractions with controlled particle-size and functionality.
- High-moisture extrusion & texturisation – twin-screw extruders configurable from 5 kg h-¹ R&D runs up to 100 kg h-¹ pilots create fibrous or puffed matrices while capturing torque/viscosity data for model calibration.
- Drying & encapsulation – lab and pilot spray-, freeze- and vacuum-dryers (1 – 30 kg h-¹) produce micro-encapsulated flavours, emulsions and shelf-stable powders ready for blending.
- Shelf-life, packaging & compliance – modified-atmosphere test chambers, migration/oxidation assays and AOAC microbiology suites deliver regulatory dossiers and commercial-ready packaging specs.
Food Technologies: Analytics
From single-cell metabolic fingerprints to consumer-perceived mouthfeel, our integrated food-tech analytics platform compresses months of iteration into days.
Purpose-built to serve the entire design-make-test-scale pipeline, it links upstream bioprocess monitoring (real-time Raman/FTIR, online metabolomics and spent-media flux analysis) with downstream compositional, safety and sensory characterisation through a secure digital-twin backbone.
When our new reference laboratory comes online in Q3 2025, we will be one of only a handful of centres accredited by the Periodic Table of Food Initiative (PTFI), ensuring every dataset—from 2,500-analyte metabolite panels to ISO-17025 microbial assays—is globally comparable and openly sharable.
Core Research Areas
- Periodic Table of Food Initiative – LC-MS, GC-MS, NMR and HR-MS workflows calibrated to PTFI protocols quantify >2 500 metabolites, bioactives and trace elements per sample.
- Spent-media insight engine – High-resolution Agilent suite chromatography, Q-TOF and ICP-MS platforms inside a dedicated measurement facility deliver untargeted targeted metabolomics and lipidomics with sub-ppm mass accuracy. tracks amino acids, vitamins and polar metabolites in fermentation broths in <10 min, guiding dynamic feed strategies.
- Ramanomics fingerprinting – high-content micro-Raman microscopy produces label-free molecular maps at single-cell or micro-domain resolution, enabling rapid discrimination of proteins, lipids and carbohydrates in complex food matrices.
- Safety & compliance toolkit – ultra-low-cost spoilage-gas sensors, rapid lateral-flow pathogen tests, and ISO microbiology panels provide real-time shelf-life, allergen and contaminant assurance.
- Texture & rheology laboratory – dynamic shear rheometers, TA.XT texture analysers and tribology rigs characterise viscoelasticity, fracture and mouth-feel, feeding data directly into digital-twin process models.


Artificial Intelligence
Artificial intelligence sits at the heart of the Centre’s “design-make-test-learn” loop: anchored by I-X, Imperial’s flagship AI hub, we are building the first end-to-end data backbone for alternative proteins—linking multi-omics, bioprocess, sensory and LCA streams into standardised, FAIR-compliant datasets ready for open science.
This infrastructure couples generative-AI strain design, AlphaFold-guided protein-function prediction and real-time digital-twin optimisation, turning weeks of wet-lab iteration into hours of in-silico exploration.
Core Research Areas
- Data lake & curation – automated pipelines gather, clean and harmonise multi-omics, process analytics technologies (PAT) and sensory data into a cloud SQL/Parquet repository.
- Generative-AI strain & enzyme design – graph neural networks propose edits; active-learning DoE closes the loop with the biofoundry to converge on high-performance variants.
- Feedstock-aware bioprocess models – hybrid physics/ML algorithms predict metabolism and yield shifts for variable waste-derived substrates, guiding adaptive control recipes.
- Taste-and-nutrition prediction – multimodal transformers link molecular fingerprints to sensory panel scores and NutriScore metrics, enabling “flavour-in-silico” screening.
- Real-time digital twins – edge-deployed ML models ingest Raman/FTIR, capacitance and off-gas signals to keep critical quality attributes within spec during scale-up.
- Open model & API portal – Git-based registry and Swagger APIs let academics or start-ups query trained models, upload new data and benchmark performance in minutes.
Nutrition
From petri-dish to personalised diet—we have capacity to test at every stage, from in-vitro digestion models to first-in-human testing, with our clinical nutrition pipeline.
The same platform underpins our new role as a Periodic Table of Food Initiative (PTFI) reference lab, so compositional data generated internally feeds directly into global open databases.
Core Research Areas
- Gastro-intestinal simulation – INFOGEST-compliant static and dynamic digestors reproduce mouth-to-ileum conditions, yielding bio-accessible fractions for rapid screening.
- Whole-body metabolic phenotyping – Stable-isotope tracers, indirect calorimetry and breath analysis map post-prandial fluxes with clinic-grade precision for both acute and longitudinal studies.
- Clinical intervention pipeline – Dedicated metabolic wards integrate dual-energy X-ray absorptiometry (DEXA), continuous glucose monitoring and genomics/microbiome sequencing to deliver Phase 0-II trials and personalised nutrition read-outs.
- Gut microbiome – impact of novel food on microbiome health

Cell banks
Spanning seeds to stem cells—anchor our platform: registry links plant, microbial, algal and mammalian stocks to full-stack omics fingerprints, so users can request, trace and deploy high-quality cell lines/ germplasms for your research fast.
Core Research Areas
- Yeast & filamentous fungi: flavour-building non-Saccharomyces, Yarrowia lipolytica, high-lipase dairy strains, robust
- GRAS bacterial chassis library: modular plasmid toolkit and genome-refactored hosts engineered for precision-fermentation of proteins, lipids and flavours.
- Immortal mammalian cell lines: bovine, avian and fish stem-cell banks developed under serum-free protocols, delivering scalable starters for cultivated-meat R&D.
- Integrated QC & traceability: barcoded cryo-vials, routine STR/ITS sequencing and cloud LIMS ensure chain-of-custody integrity across every shipment.